In The News
27 May 2026
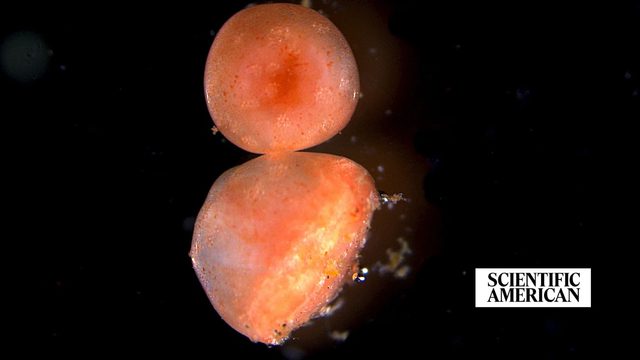
Amputated sea cucumber tissue keeps living for years—possibly forever
From Scientific American, President Alejandro Sánchez Alvarado, Ph.D., shares expert insight on a fascinating new regeneration study.
Read Article
By Anissa Orr
In a perfect world geared toward advancing science, researchers would have the resources to explore whatever promising ideas came their way. But the funding process, which often rewards sure things instead of unproven concepts, forces many scientists to put their riskier, and perhaps best, ideas on the backburner.
Thankfully that's not the case for Stowers Associate Investigators Ron Yu, PhD, and Julia Zeitlinger, PhD. Both are recipients of the 2015 Neaves Award, established in honor of William B. Neaves, PhD, president emeritus of the Stowers Institute for Medical Research. The two-year, $150,000 award was launched in 2010 to encourage and support Stowers researchers who wish to pursue innovative, high-risk research projects with the potential for broad impact.
Clearer, faster images of the nervous system
Yu's work focuses on the sensory system and identifying the complex connections between neurons that link sensation to behavior. "The mammalian main olfactory and vomeronasal systems give us two ways to look at it," he says.
Yu explains that the olfactory system generally triggers learned behaviors. The smell of freshlybaked chocolate chip cookies evokes happy childhood memories of cooking with a loved one, while the pungent smell of disinfectant calls up unhappy memories of a hospital stay. The triggered memory may influence our behavior (either reaching for a cookie, or fleeing the hospital). The associations we make with smells are often learned, Yu emphasizes.
In contrast, sensory input in the vomeronasal system in mice triggers involuntary or innate behaviors such as the mating process. Yu's 2014 study on this system published in the leading science journal eLife, offered new insight into how the mammalian brain processed pheromones to elicit courtship behaviors.
Yu will use the Neaves Award to develop a new tool that adds compressed sensing technology to microscopes, enabling rapid imaging of neuronal circuits in the brain. "The idea is to create a device that allows a microscope with the same number of sensors to pick up more signals from our biological sample to increase either the resolution of the image or the speed of acquisition," Yu says. If successful, he expects this tool to become a fully integral part of microscopy systems. Yu plans to use the technology to image the entire mouse brain and trace its neurocircuitry.
Currently, Yu and his team are working on a prototype using software developed by Dar Dahlen, a bioinformatician formerly in the lab. Enlisting scientific advisors and collaborators inside and outside of the Institute, Yu's team will test and implement several designs of the tool's hardware component, which could be attached to a microscope to improve resolution. The funds will also allow Yu to hire another research associate dedicated to the project.
Yu says he never imagined when he joined the Institute in 2004 that he would one day help create a new type of microscopy technology, but the Neaves Award has helped him step out of his comfort zone to pursue research wherever it takes him. "I don't think this project would have been approved by traditional funding mechanisms," he says. "With the Neaves Award, we have the freedom to explore."
Innovative improvements in DNA mapping
The desire to uncover the rules that govern gene regulation drives Zeitlinger's research, and has been her passion since she joined the Institute in 2007. By investigating the general principles behind gene regulation, Zeitlinger hopes to gain insight that can be applied to the human genome in development and disease.
Gene regulation is the process by which a cell determines which genes to turn on and off and when to create specific cells in the body, for example, a neuron, a brain cell, or the cells that make up skin or bone.
This process usually happens during transcription, when the information in a gene's DNA is transcribed or transferred to RNA, and involves proteins called transcription factors. Transcription factors bind to specific spots in the DNA to switch on and off certain genes. Researchers know that these binding sites are important, but still have trouble finding their exact location.
Determined to zero in on these areas, Zeitlinger and her team developed innovative improvements to a technique called ChIP-seq, which gives investigators an accurate high-resolution map of transcription binding sites.
She and colleagues Qiye He, PhD, and Jeff Johnston published their first paper on the improved technique, called ChIP-nexus, in the April 2015 issue of Nature Biotechnology. ChIP-nexus is already yielding new information, showing that binding factors are not scattered across the genome as previously thought, but rather appear in specific, predictable sequences.
Now Zeitlinger is using the Neaves Award to make further tweaks to ChIP-seq technology that will allow researchers to analyze transcription factor binding in populations of cells too small to be characterized by previous technologies. Zeitlinger believes the resulting novel technique, deemed ChIP-next, could be a game changer for genetic researchers, particularly those who prefer to work with cells from intact tissue, which can be limited in quantity, instead of more abundant cultured cells.
Intact tissue cells are often preferred over cultured cells because they can be examined in their natural context. However, using current technologies, a vast amount of live cells—sometimes numbering in the millions—need to be collected per experiment to provide an accurate picture of the genome. "If you want to study gene regulation in real organisms, from embryos or specific tissues in the body, you don't have that many cells available," Zeitlinger says. "ChIP-next is ideal for researchers investigating a specific stem cell niche or a particular system in the body."
With funds from the Neaves Award, she plans to implement ChIPnext and add another researcher to the project. Zeitlinger credits the award for continuing the project's momentum.
"If this technology can be used for much more specific cell types, then it could be attractive to many scientists and a great deal of regulatory information could be collected over time. This would have a huge impact on the field," Zeitlinger says, adding that the technique also holds promise for human health. At the moment, linking genes with particular developmental defects is difficult at best. "But, having all this information will make it much easier to make associations between a developmental defect and what went off course genetically."
Continuing the legacy of Dr. William B. Neaves
Both researchers' projects exemplify the type of innovative research the Neaves award was established to support, says the award's namesake, William (Bill) B. Neaves, PhD. Neaves served as Stowers' President and CEO from 2000 to 2010, recruiting world-class scientists to the then-fledgling institution and fostering an environment where researchers and their work flourished. He's happy to see his legacy continue with the Neaves Award.
"I cherish my association with the outstanding research programs of Ron Yu and Julia Zeitlinger," Neaves says. "Together with Robb Krumlauf, I had the privilege of recruiting Ron from Columbia University and Julia from Massachusetts Institute of Technology. Now that each of them has received the award established by the Stowers Institute in my name, I feel doubly honored."
Stowers President and CEO Dave Chao, PhD, explains that Neaves played an invaluable role in creating an environment that encourages its researchers to pursue challenging questions with novel and creative approaches. "For more than a decade, Bill has always embodied and fostered a pioneering spirit at the Institute," he says, "An award enabling Stowers investigators to pursue creative and ambitious work in new areas is aptly named to honor Bill."
In The News

27 May 2026
From Scientific American, President Alejandro Sánchez Alvarado, Ph.D., shares expert insight on a fascinating new regeneration study.
Read Article
In The News

22 May 2026
Former Stowers Graduate School Summer Scholar Isaac Witte, Ph.D., was featured in the Harvard Gazette ahead of his graduation this month.
Read Article
News
19 May 2026
For Stowers Investigator Linheng Li, Ph.D., a new leukemia study builds on a career spent asking how the places stem cells call home can shape health, disease, and future treatments
Read Article
